In the recent Critical Assessment of Protein Structure Prediction 5 (CASP13)-a blind assessment of the state of the field-AlphaFold created high-accuracy structures (with template modelling (TM) scores 6 of 0.7 or higher) for 24 out of 43 free modelling domains, whereas the next best method, which used sampling and contact information, achieved such accuracy for only 14 out of 43 domains. The resulting system, named AlphaFold, achieves high accuracy, even for sequences with fewer homologous sequences. We find that the resulting potential can be optimized by a simple gradient descent algorithm to generate structures without complex sampling procedures. 
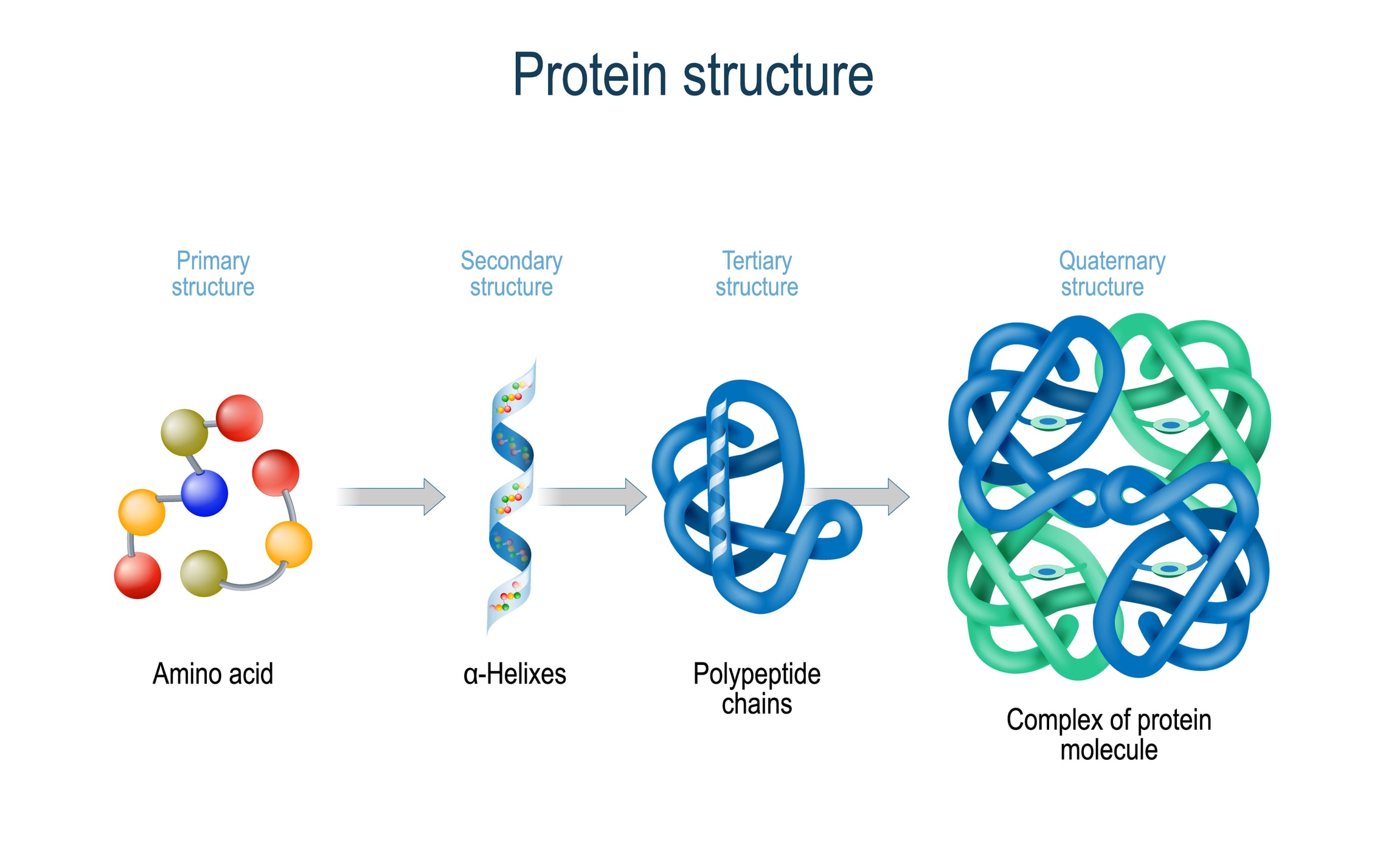
Using this information, we construct a potential of mean force 4 that can accurately describe the shape of a protein. Here we show that we can train a neural network to make accurate predictions of the distances between pairs of residues, which convey more information about the structure than contact predictions. It is possible to infer which amino acid residues are in contact by analysing covariation in homologous sequences, which aids in the prediction of protein structures 3. Considerable progress has recently been made by leveraging genetic information. This problem is of fundamental importance as the structure of a protein largely determines its function 2 however, protein structures can be difficult to determine experimentally. Protein structure prediction can be used to determine the three-dimensional shape of a protein from its amino acid sequence 1.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
